urdf to collada for OpenRave does not work
I am using OpenRave for UR5e fastIK calculation. Since I need to load collada file, I am converting the ur5e xacro files in ur_e_description package by following these steps:
xacro to urdf:
xacro ur5e_robot.urdf.xacro >urdf to collada:
rosrun collada_urdf urdf_to_collada ur5e.urdf ur5e.dae
And then I use this file in the OpenRave simulation. The result is like that: 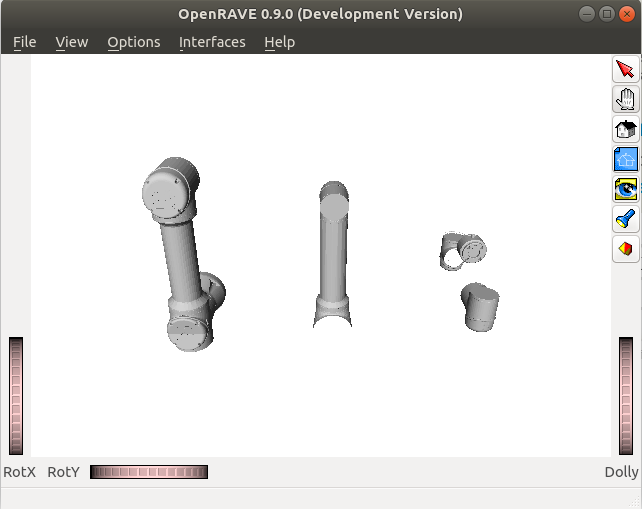
The calculated end effector pose looks right, though.
Tee_pose:
position: x: 0.8171999999999516 y: 0.23289999995927044 z: 0.0627999999522314
orientation: x: 0.7071067811865479 y: 0.707106781186547 z: -6.905298555182071e-11 w: 7.266121038185247e-11
I have spent lots of time figuring out why such a thing happens; is it during the conversion from xacro to urdf or the urdf into collada. Then I tried to load the produced collada (ur5e.dae) into moveit_setup_assistant, it loads without any issue: 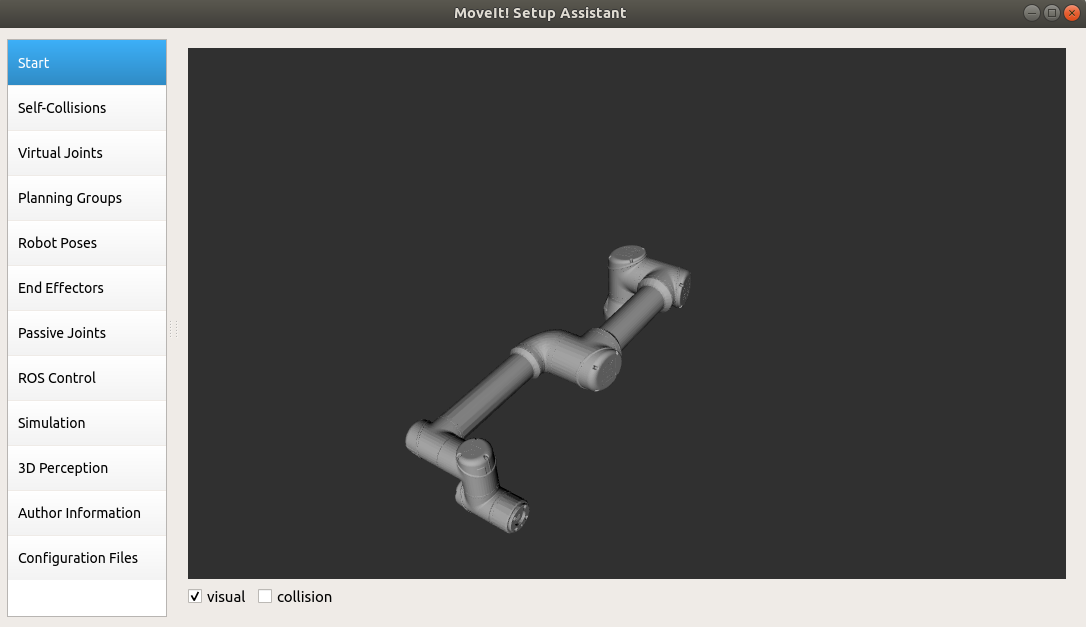 .
.
Interestingly, I can manipulate the robot in OpenRave by setting DOF values. However, I cannot produce any ikmodel since (I suppose) the produced urdf/collada file is broken. When I check the produced urdf (from xacro) I saw some interesting values. For instance for shoulder_pan_joint the DH parameters are like that: 
Yet the corresponding part of the urdf, there is pi rotation in yaw direction (not pi/2):
<joint name="shoulder_pan_joint" type="revolute">
<parent link="base_link"/>
<child link="shoulder_link"/>
<origin rpy="0 0 3.14159265359" xyz="0 0 0.1625"/>
<axis xyz="0 0 1"/>
<limit effort="150.0" lower="-6.28318530718" upper="6.28318530718" velocity="3.14"/>
<dynamics damping="0.0" friction="0.0"/>
</joint>
Does anyone know why such behaviour occur? What is it about and how to solve it?


Having same issue, any luck?
Yes and no, I haven't solved this specific problem but I loaded another urdf file (not directly from Universal Robots ROS package). My guess is that some parametrized values or included xacros are not properly loaded in OpenRAVE, but I am not sure. The working urdf does not contain those complex features.
If you just want to generate IK solver C++ output file then I will suggest to use your generated collada only since it is correct and only throughs warning which are due to some incorrect transpency values set during the conversion.
No, I also needed to simulate in openrave. Therefore, I changed into a non-parametrized urdf.